Expert genomics analysis services for all major sequencing platforms at Symba Genomics
Modern genomics research demands the right sequencing technology for optimal results. Illumina NovaSeq, PacBio HiFi, and Oxford Nanopore each offer distinct advantages for different applications. This guide provides essential comparisons to help researchers select the optimal platform for their projects. The three platforms serve different niches in the genomics ecosystem. Illumina dominates high-throughput applications, PacBio excels at accurate long-read assembly, while Oxford Nanopore enables real-time sequencing with ultra-long reads. This summary table highlights the key differentiating factors across all three platforms, enabling quick comparison of technical specifications and practical considerations for project planning
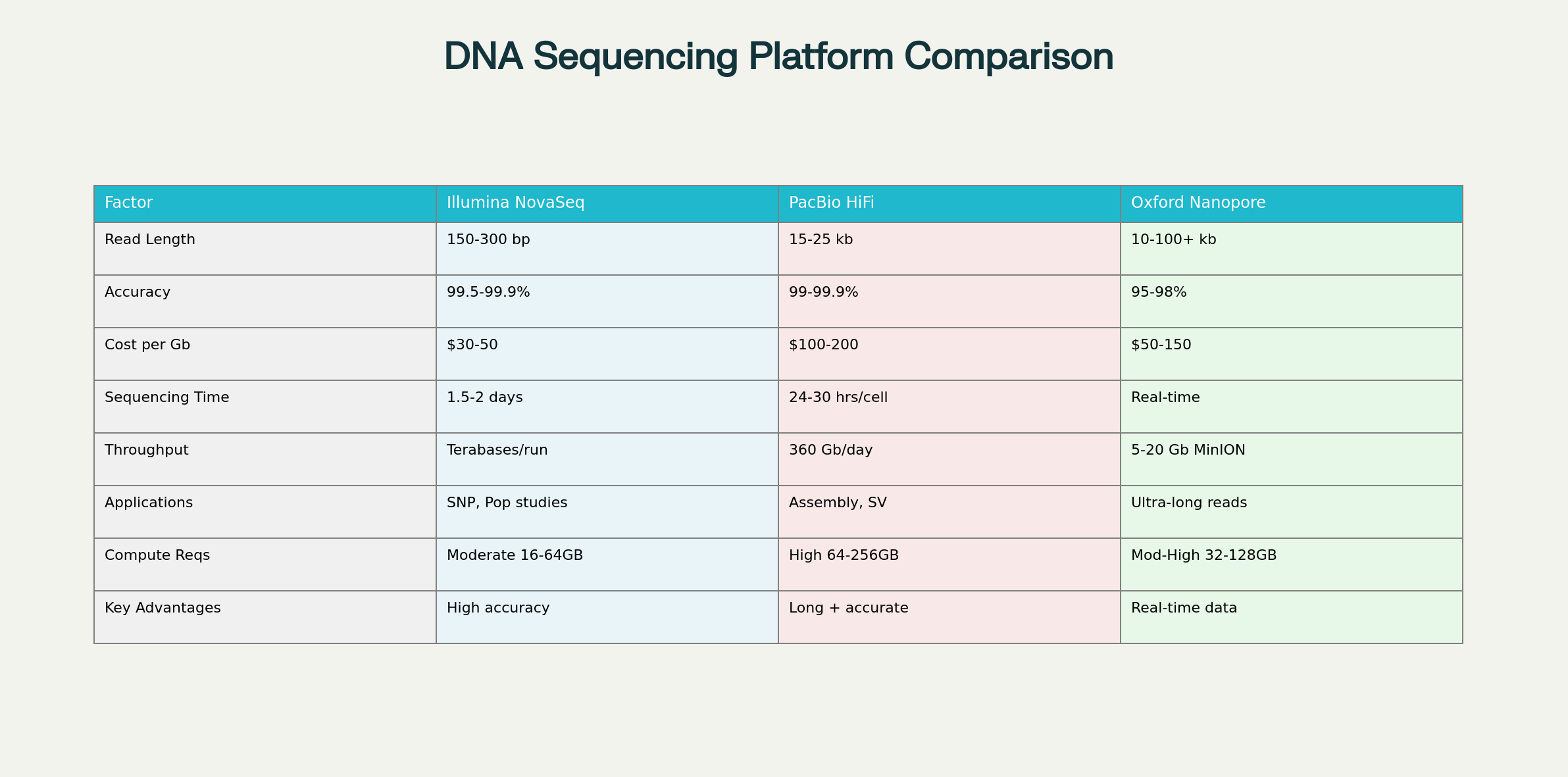
Key Technical Differences
Read Length Capabilities:
The dramatic differences in read length capabilities fundamentally determine each platform’s applications. Short reads excel at SNP detection but cannot span repetitive regions, while long reads enable complete genome assembly and structural variant detection.
Cost vs Performance Analysis
Illumina NovaSeq offers the most cost-effective solution for high-accuracy applications, making it ideal for population studies and clinical diagnostics. PacBio HiFi commands premium pricing but delivers unmatched accuracy for long-read applications. Oxford Nanopore provides balanced cost-performance with unique real-time capabilities.
Bioinformatics Considerations: Tools and Workflows
The computational requirements and analysis pipelines differ significantly between platforms. Illumina benefits from decades of tool development, with mature, standardized pipelines for alignment (BWA-MEM, Bowtie2) and variant calling (GATK, FreeBayes). These workflows are highly optimized and widely adopted across research and clinical communities. Long-read analysis requires more specialized approaches. Modern tools like minimap2 for alignment and assemblers like Hifiasm have matured considerably, but workflows remain more flexible and require greater expertise. PacBio HiFi data, given its accuracy, integrates well with existing variant calling tools, while Oxford Nanopore data often requires additional basecalling steps and may benefit from error correction strategies. Computational resource requirements vary by platform. While Illumina generates massive read numbers requiring substantial storage and processing power, long-read platforms produce fewer reads but may require high-memory servers for genome assembly tasks. Oxford Nanopore’s basecalling process can be computationally intensive, often utilizing GPUs for real-time processing.
Application-Specific Platform Selection
Clinical Application
Illumina sequencing remains the clinical standard for routine diagnostics due to its validated accuracy, but it struggles with detecting large structural variants and repeat expansions that cause certain genetic diseases. PacBio HiFi is emerging as a powerful complement for complex cases where Illumina yields inconclusive results, offering the ability to resolve structural variants and phase haplotypes that may reveal previously missed pathogenic variants. Oxford Nanopore has established a unique role in rapid diagnostics, enabling 8-hour whole-genome sequencing for critically ill patients and portable pathogen sequencing during disease outbreaks.
Research applications
In metagenomic research, Illumina excels at community profiling and quantitative analysis but struggles with assembling complete microbial genomes, while long-read platforms like PacBio HiFi and Oxford Nanopore can assemble bacterial genomes into single contiguous sequences, including repetitive elements that short reads cannot resolve. For whole-genome sequencing, Illumina remains ideal for population studies focused on small variant detection, but long-read platforms are essential for de novo assembly and structural variant analysis. PacBio HiFi has become the preferred choice for reference-grade genome assembly with exceptional accuracy, while Oxford Nanopore’s ultra-long reads enable complete telomere-to-telomere assemblies by spanning the largest repetitive elements.
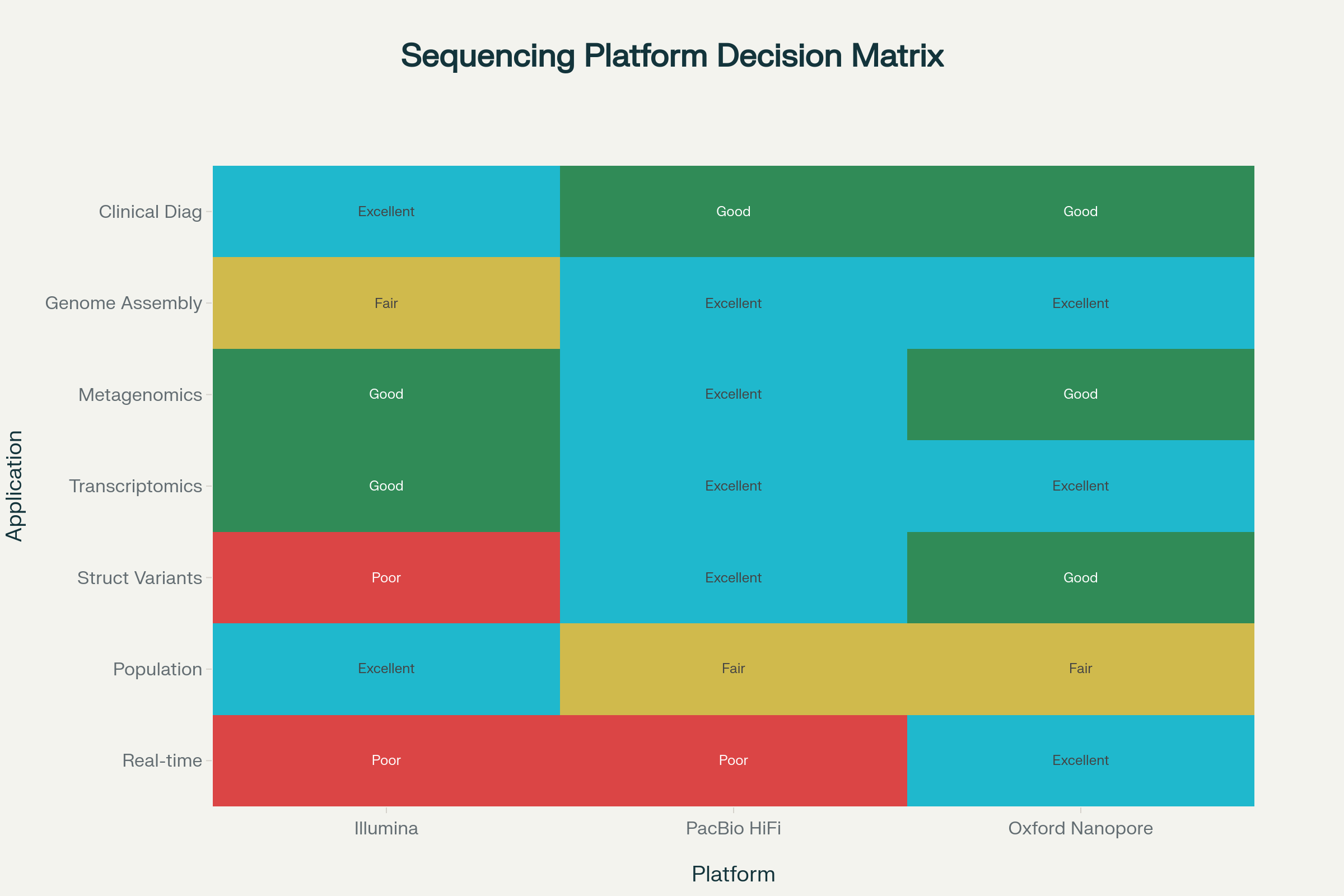
Platform suitability matrix for genomics applications
Data Processing Workflows
Illumina:
Established pipelines with extensive automation and quality control standards.
PacBio HiFi:
Consensus-based workflows requiring specialized assembly and polishing steps.
Oxford Nanopore:
Real-time basecalling with rapidly evolving analysis methods and direct modification detection.
Platform Selection Recommendation
| Illumina Novaseq | PacBio HiFi | Oxford Nanopore |
|---|
- Population studies (1000+ samples)
- Clinical diagnostics and gene panels
- SNP association studies
- Cost-sensitive large-scale projects
| - Reference-quality genome assemblies
- Structural variant analysis
- Complex genetic disorders
- High-accuracy long-read requirements
| - Rapid diagnostic sequencing
- Field-based genomics applications
- Methylation and epigenetic studies
- Ultra-long read assembly projects
|
Conclusion
Platform selection depends on
accuracy requirements, read length needs, timeline constraints, and budget considerations. Rather than competing technologies, these platforms function as complementary tools in modern genomics.
Illumina remains optimal for high-throughput, cost-sensitive applications with exceptional accuracy.
PacBio HiFi excels where long-read accuracy is essential for complex genomic analysis.
Oxford Nanopore provides unique real-time capabilities and ultra-long reads for specialized applications. At
Symba Genomics, our multi-platform expertise ensures optimal technology selection and analysis strategies for your research objectives. Whether you need comprehensive genome assembly, clinical diagnostic support, or population-scale analysis, we deliver expert services tailored to your requirements.
Expert Analysis Services at Symba Genomics
Comprehensive Platform Support
Pre-Project Consultation:
- Platform selection based on research objectives
- Cost-benefit analysis and timeline planning
- Sample preparation guidance
- Quality assessment protocols
Multi-Platform Integration:
- Hybrid assembly strategies
- Cross-platform validation
- Comprehensive variant annotation
Contact us to discuss your genomics project and discover how our platform expertise accelerates research success.

 Platform suitability matrix for genomics applications
Platform suitability matrix for genomics applications